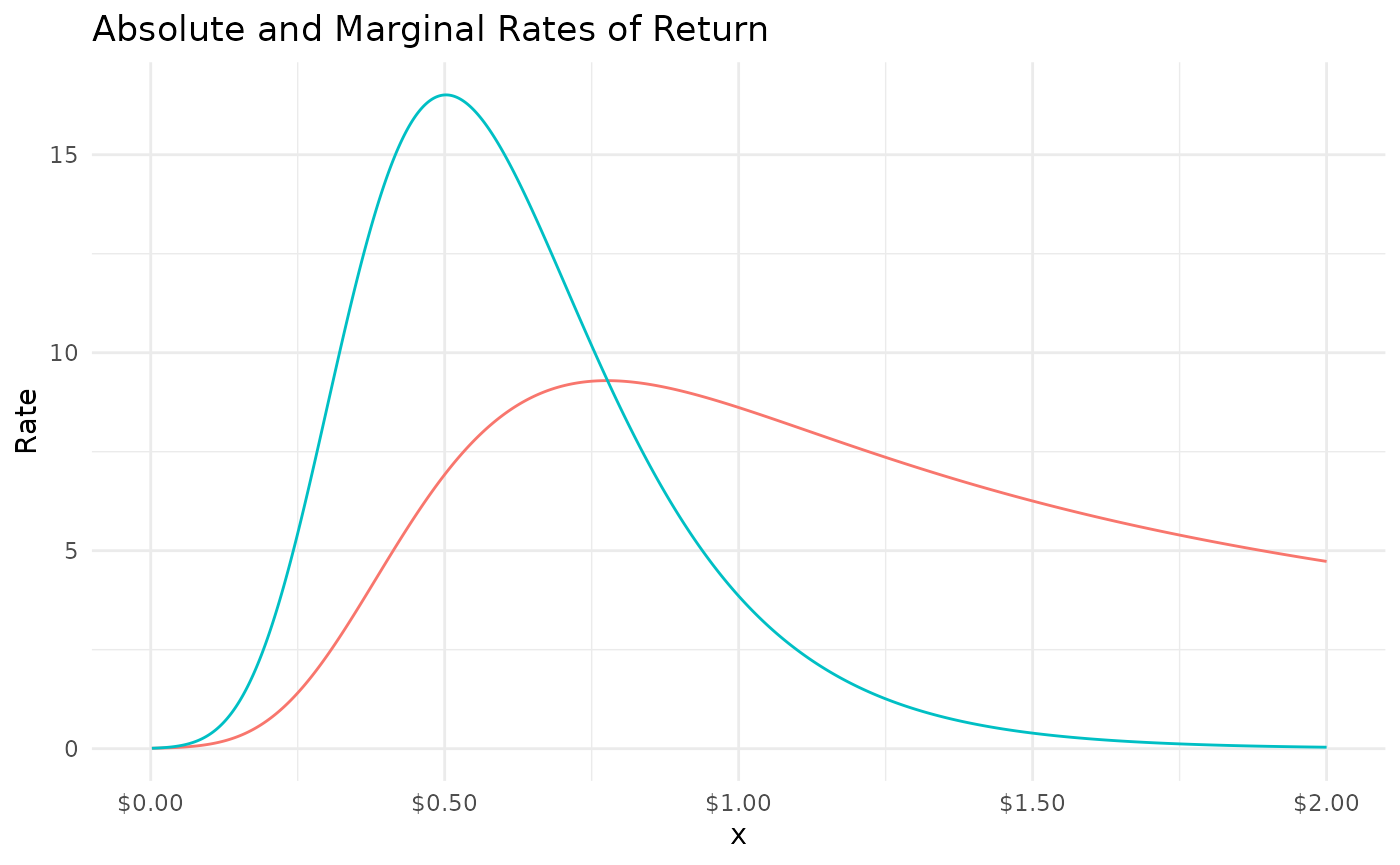
library(mrmopt)
library(brms)
#> Loading required package: Rcpp
#> Loading 'brms' package (version 2.23.0). Useful instructions
#> can be found by typing help('brms'). A more detailed introduction
#> to the package is available through vignette('brms_overview').
#>
#> Attaching package: 'brms'
#> The following object is masked from 'package:stats':
#>
#> ar
library(purrr)
library(gt)
library(dplyr)
#>
#> Attaching package: 'dplyr'
#> The following objects are masked from 'package:stats':
#>
#> filter, lag
#> The following objects are masked from 'package:base':
#>
#> intersect, setdiff, setequal, union
library(ggplot2)
library(patchwork)
x_values <- seq(0, 1, by = 0.01)
b <- -5
c <- 1
d <- 10
e <- 0.5
y_response = rm_Gompertz(x_values, b, c, d, e) + rnorm(length(x_values), mean = 0, sd = 0.4)
response_df <- bind_cols(
x = x_values,
y = y_response
)
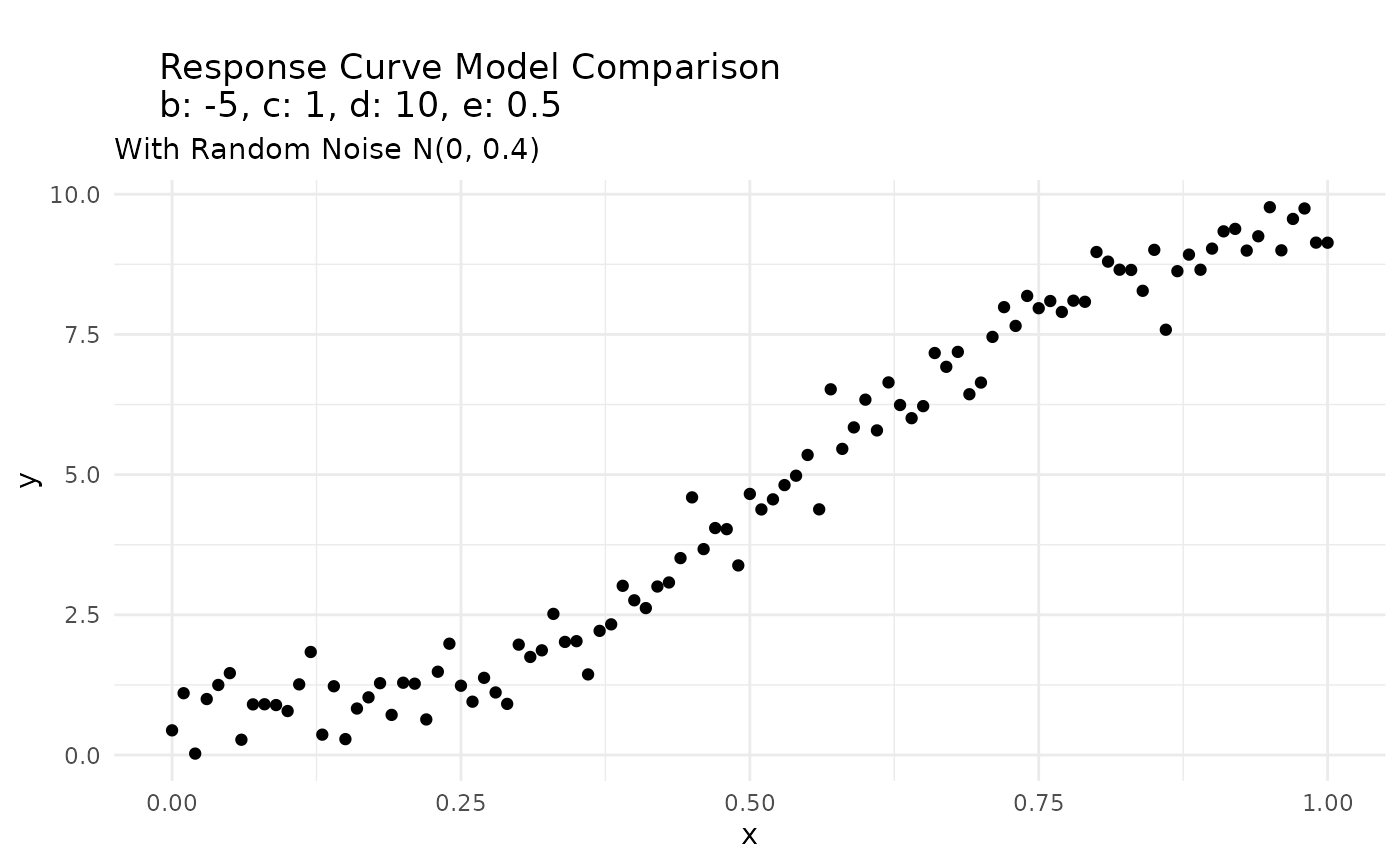
response_df |>
ggplot(aes(x, y)) +
geom_point() +
labs(title = paste0("
Response Curve Model Comparison
", "b: ", b, ", c: ", c, ", d: ", d, ", e: ", e),
subtitle = "With Random Noise N(0, 0.4)"
) +
theme_minimal()
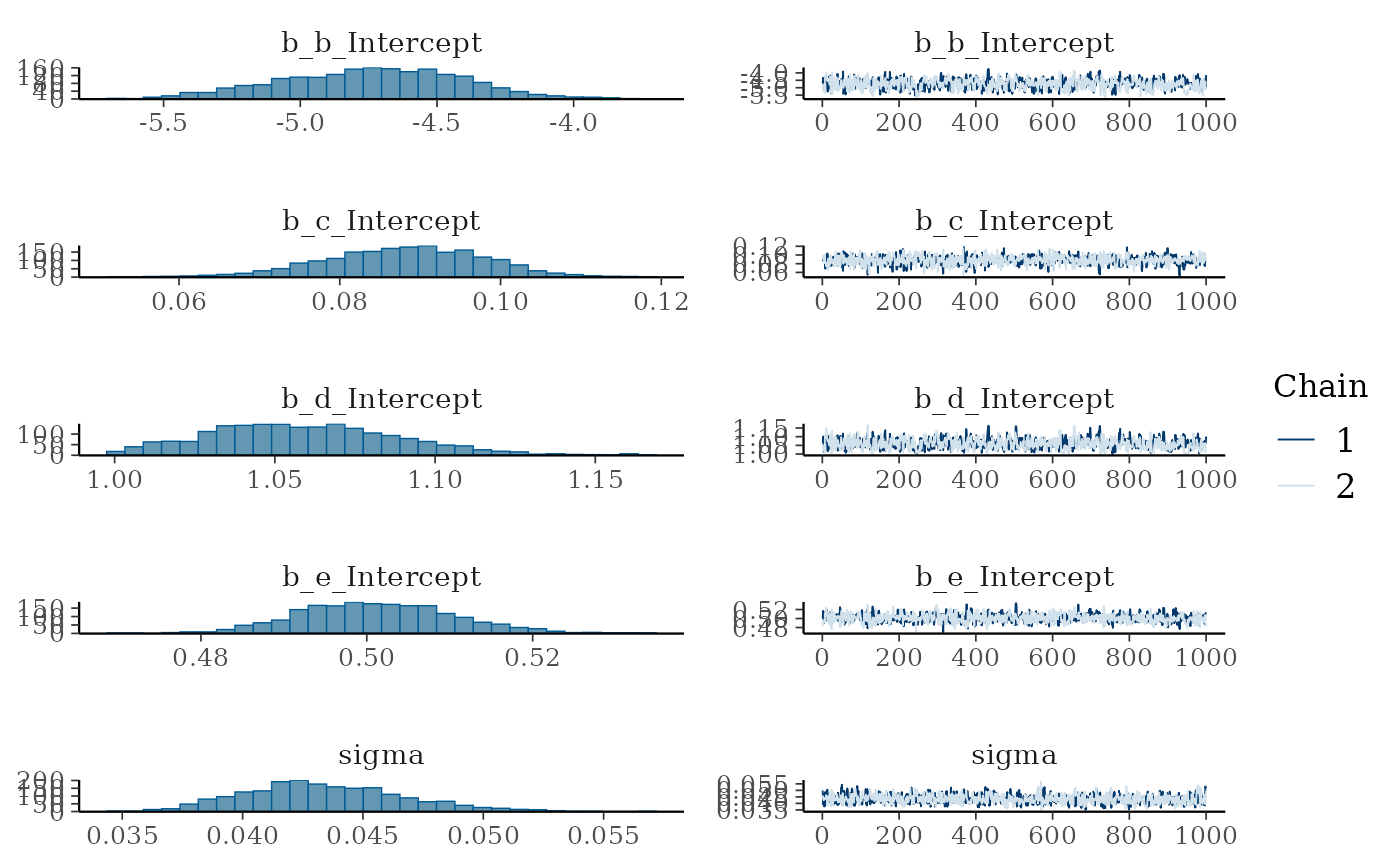
response_fit =
fit_response(
data = response_df,
x = "x",
y = "y",
type = "gompertz",
auto = TRUE,
# prior = c(
# prior(normal(-5, 10), nlpar = "b", lb = -10, ub = 0),
# prior(normal(0, 5), nlpar = "c", ub = 4.9),
# prior(normal(10, 5), nlpar = "d", lb = 5.1),
# prior(normal(0.6, 2), nlpar = "e")
# ),
chains = 2,
iter = 2000,
warmup = 1000,
seed = 007,
control = list(adapt_delta = 0.90)
)
#> y ~ c + (d - c) * exp(-exp(b * (x - e)))
#> b ~ 1
#> c ~ 1
#> d ~ 1
#> e ~ 1
#>
#> SAMPLING FOR MODEL 'anon_model' NOW (CHAIN 1).
#> Chain 1:
#> Chain 1: Gradient evaluation took 0.000126 seconds
#> Chain 1: 1000 transitions using 10 leapfrog steps per transition would take 1.26 seconds.
#> Chain 1: Adjust your expectations accordingly!
#> Chain 1:
#> Chain 1:
#> Chain 1: Iteration: 1 / 2000 [ 0%] (Warmup)
#> Chain 1: Iteration: 200 / 2000 [ 10%] (Warmup)
#> Chain 1: Iteration: 400 / 2000 [ 20%] (Warmup)
#> Chain 1: Iteration: 600 / 2000 [ 30%] (Warmup)
#> Chain 1: Iteration: 800 / 2000 [ 40%] (Warmup)
#> Chain 1: Iteration: 1000 / 2000 [ 50%] (Warmup)
#> Chain 1: Iteration: 1001 / 2000 [ 50%] (Sampling)
#> Chain 1: Iteration: 1200 / 2000 [ 60%] (Sampling)
#> Chain 1: Iteration: 1400 / 2000 [ 70%] (Sampling)
#> Chain 1: Iteration: 1600 / 2000 [ 80%] (Sampling)
#> Chain 1: Iteration: 1800 / 2000 [ 90%] (Sampling)
#> Chain 1: Iteration: 2000 / 2000 [100%] (Sampling)
#> Chain 1:
#> Chain 1: Elapsed Time: 1.04 seconds (Warm-up)
#> Chain 1: 1.056 seconds (Sampling)
#> Chain 1: 2.096 seconds (Total)
#> Chain 1:
#>
#> SAMPLING FOR MODEL 'anon_model' NOW (CHAIN 2).
#> Chain 2:
#> Chain 2: Gradient evaluation took 7.9e-05 seconds
#> Chain 2: 1000 transitions using 10 leapfrog steps per transition would take 0.79 seconds.
#> Chain 2: Adjust your expectations accordingly!
#> Chain 2:
#> Chain 2:
#> Chain 2: Iteration: 1 / 2000 [ 0%] (Warmup)
#> Chain 2: Iteration: 200 / 2000 [ 10%] (Warmup)
#> Chain 2: Iteration: 400 / 2000 [ 20%] (Warmup)
#> Chain 2: Iteration: 600 / 2000 [ 30%] (Warmup)
#> Chain 2: Iteration: 800 / 2000 [ 40%] (Warmup)
#> Chain 2: Iteration: 1000 / 2000 [ 50%] (Warmup)
#> Chain 2: Iteration: 1001 / 2000 [ 50%] (Sampling)
#> Chain 2: Iteration: 1200 / 2000 [ 60%] (Sampling)
#> Chain 2: Iteration: 1400 / 2000 [ 70%] (Sampling)
#> Chain 2: Iteration: 1600 / 2000 [ 80%] (Sampling)
#> Chain 2: Iteration: 1800 / 2000 [ 90%] (Sampling)
#> Chain 2: Iteration: 2000 / 2000 [100%] (Sampling)
#> Chain 2:
#> Chain 2: Elapsed Time: 1.284 seconds (Warm-up)
#> Chain 2: 1.024 seconds (Sampling)
#> Chain 2: 2.308 seconds (Total)
#> Chain 2:
summary(response_fit)
#> Family: gaussian
#> Links: mu = identity
#> Formula: y ~ c + (d - c) * exp(-exp(b * (x - e)))
#> b ~ 1
#> c ~ 1
#> d ~ 1
#> e ~ 1
#> Data: data (Number of observations: 101)
#> Draws: 2 chains, each with iter = 2000; warmup = 1000; thin = 1;
#> total post-warmup draws = 2000
#>
#> Regression Coefficients:
#> Estimate Est.Error l-95% CI u-95% CI Rhat Bulk_ESS Tail_ESS
#> b_Intercept -4.74 0.33 -5.40 -4.12 1.01 576 794
#> c_Intercept 0.09 0.01 0.07 0.11 1.01 950 1090
#> d_Intercept 1.06 0.03 1.01 1.12 1.01 542 435
#> e_Intercept 0.50 0.01 0.48 0.52 1.00 924 943
#>
#> Further Distributional Parameters:
#> Estimate Est.Error l-95% CI u-95% CI Rhat Bulk_ESS Tail_ESS
#> sigma 0.04 0.00 0.04 0.05 1.00 894 842
#>
#> Draws were sampled using sampling(NUTS). For each parameter, Bulk_ESS
#> and Tail_ESS are effective sample size measures, and Rhat is the potential
#> scale reduction factor on split chains (at convergence, Rhat = 1).
| Parameter Estimates vs Actual Values |
| Using the Gompertz Response Model |
| actual |
center |
lower |
upper |
| -5.0 |
-4.74 |
-5.40 |
-4.12 |
| 1.0 |
0.88 |
0.68 |
1.06 |
| 10.0 |
10.35 |
9.85 |
10.95 |
| 0.5 |
0.50 |
0.48 |
0.52 |
mrm_params_table(response_fit, scaled = FALSE) |>
gt::tab_header(
title = "Parameter Estimates Table",
subtitle = "Using the Gompertz Response Model"
)
| Parameter Estimates Table |
| Using the Gompertz Response Model |
| Parameter |
center |
lower |
upper |
| b (growth rate) |
-4.741645 |
-5.398366 |
-4.123516 |
| c (lower asymptote) |
0.08800686 |
0.06760201 |
0.1062646 |
| d (upper asymptote) |
1.059448 |
1.008309 |
1.120964 |
| e (inflection point) |
0.5012417 |
0.4843928 |
0.5204534 |
response = mrm_infer(response_fit, length.out = 100)
head(response) |>
gt::gt() |>
gt::tab_header(
title = "Inferred Response Data",
subtitle = "Using the Gompertz Response Model"
) |>
gt::fmt_number(
columns = tidyselect::everything(),
decimals = 2
)
| Inferred Response Data |
| Using the Gompertz Response Model |
| x |
Estimate |
Est.Error |
Q2.5 |
Q97.5 |
center |
lower |
upper |
type |
resp_var |
input_var |
ar |
mr |
cp |
cp_lower |
ar_lower |
mr_lower |
ar_upper |
mr_upper |
cp_upper |
| 0.00 |
0.89 |
0.43 |
0.07 |
1.73 |
0.88 |
0.03 |
1.74 |
gompertz |
y |
x |
NaN |
NA |
NaN |
NaN |
NaN |
NA |
NaN |
NA |
NaN |
| 0.02 |
0.89 |
0.43 |
0.00 |
1.77 |
0.88 |
0.03 |
1.74 |
gompertz |
y |
x |
0.02 |
0.02 |
53.35 |
60.60 |
0.03 |
0.03 |
0.00 |
0.00 |
3,718.16 |
| 0.04 |
0.88 |
0.43 |
0.04 |
1.74 |
0.88 |
0.03 |
1.74 |
gompertz |
y |
x |
0.03 |
0.04 |
32.26 |
36.65 |
0.04 |
0.06 |
0.01 |
0.02 |
76.30 |
| 0.06 |
0.90 |
0.43 |
0.01 |
1.74 |
0.89 |
0.03 |
1.74 |
gompertz |
y |
x |
0.05 |
0.08 |
19.52 |
22.17 |
0.06 |
0.10 |
0.03 |
0.06 |
28.98 |
| 0.08 |
0.90 |
0.43 |
0.01 |
1.74 |
0.89 |
0.04 |
1.74 |
gompertz |
y |
x |
0.07 |
0.16 |
11.99 |
13.62 |
0.09 |
0.18 |
0.05 |
0.14 |
14.49 |
| 0.10 |
0.90 |
0.43 |
0.03 |
1.79 |
0.89 |
0.04 |
1.74 |
gompertz |
y |
x |
0.12 |
0.29 |
7.54 |
8.56 |
0.14 |
0.31 |
0.10 |
0.27 |
8.17 |